Hallmarks of Aging
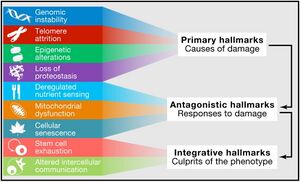
Aging is characterized by a progressive loss of physiological integrity, leading to impaired function and increased vulnerability to death. The hallmarks of aging are the types of biochemical changes that occur in all organisms that experience biological aging and lead to a progressive loss of physiological integrity, impaired function and, eventually, death. They were first listed in a landmark paper in 2013[1] to conceptualize the essence of biological aging and its underlying mechanisms.
Criteria
Each hallmark was chosen to try to fulfill the following criteria:[1]
- manifests during normal aging;
- experimentally increasing it accelerates aging;
- experimentally amending it slows the normal aging process and increases healthy lifespan.
These conditions are met to different extents by each of these hallmarks. The last criterion is not present in many of the hallmarks, as science has not yet found feasible ways to amend these problems in living organisms.
Overview
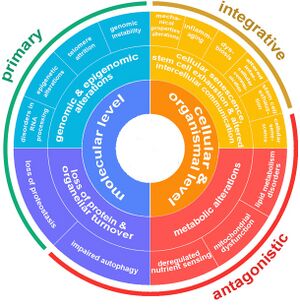
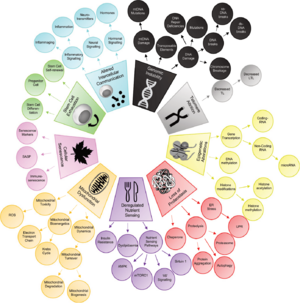
Aging is a complex process characterized by a gradual decline in physiological function. The hallmarks of aging are classified into three categories, each describing different aspects of the aging process:
- Primary Hallmarks: These are considered the main causes of cellular damage leading to aging. They are the initiating factors that, over time, drive the functional decline seen in aging cells and tissues.
- Antagonistic Hallmarks: These hallmarks are the response to the damage caused by the primary hallmarks. Initially, they may be compensatory or protective, but when chronic or excessive, they become deleterious, contributing to the aging process.
- Integrative Hallmarks: These hallmarks are the culprits of the phenotype of aging. They result from a combination of the primary and antagonistic hallmarks and are ultimately responsible for the functional decline in tissues and organs seen in aging.
The Nine Hallmarks of Aging (2013)
The nine hallmarks of aging were originally conceptualized by López-Otín and colleagues in 2013[1]. Since that it has served as a foundational paradigm for aging research for a decade until it has been revised in 2022. The original 9 landmarks were defined as follows:
| # | Hallmark | |
|---|---|---|
| 1 | Genomic Instability | Primary Hallmarks
(causes damage) |
| 2 | Telomere Attrition | |
| 3 | Epigenetic Alterations | |
| 4 | Loss of Proteostasis | |
| 5 | Deregulated Nutrient Sensing | Antagonistic Hallmarks
(responses to damage) |
| 6 | Mitochondrial Dysfunction | |
| 7 | Cellular Senescence | |
| 8 | Stem Cell Exhaustion | Integrative Hallmarks
(culprits of the phenotype) |
| 9 | Altered Intercellular Communication |
Five New Hallmarks of Aging (2022)
While these nine hallmarks have significantly advanced our understanding of aging and its relation to age-related diseases, recent critiques and evolving scientific evidence have prompted the scientific community to reconsider and expand this framework.[4] To address this, a symposium titled “New Hallmarks of Ageing” was held in Copenhagen on March 2022, where leading experts gathered to discuss potential additions and recontextualizations of these aging hallmarks. The symposium highlighted the critical need for an expanded, inclusive paradigm that encompasses newly identified processes contributing to aging. The discussions suggested the integration of five additional hallmarks:
- compromised autophagy
- dysregulation in RNA splicing
- inflammation
- loss of cytoskeleton integrity
- disturbance of the microbiome (dysbiosis)
These potential new hallmarks, along with the original nine, underscore a more comprehensive understanding of the aging process, acknowledging its multifaceted nature and its profound implications for human health and longevity.[5]
The Twelve Hallmarks of Aging (2023)
The original authors of the nine hallmarks of aging update the set of proposed hallmarks after a decade.[6]
| Level | Hallmark | Description | Proposed Year |
Category | |
|---|---|---|---|---|---|
| Molecular Level
|

|
Genomic instability | Accumulation of DNA damage over time leading to cellular dysfunction. | 2013 | Primary Hallmarks (causes damage) |
|
Telomere attrition | Reduction in the length of telomeres leading to cellular aging. | 2013 | ||

|
Epigenetic alterations | Changes in DNA methylation and histone modification affecting gene expression. | 2013 | ||

|
Loss of proteostasis | Disruption in protein folding and stability leading to cell damage. | 2013 | ||

|
Disabled macroautophagy | Impaired cellular maintenance through the consumption of own components. | 2021[7] | Antagonistic Hallmarks (responses to damage) | |
| Cellular & Organismal Level
|

|
Deregulated nutrient sensing | Alterations in nutrient sensing pathways affecting metabolism and aging. | 2013 | |
|
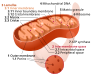
Mitochondrial dysfunction | Decrease in mitochondrial efficiency and increase in oxidative stress. | 2013 | ||

|
Cellular senescence | Accumulation of non-dividing, dysfunctional cells secreting harmful factors. | 2013 | ||
|
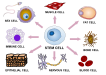
Stem cell exhaustion | Decline in the regenerative capacity of stem cells affecting tissue repair. | 2013 | Integrative Hallmarks (culprits of the phenotype) | |
|
Dysbiosis (Microbiome disturbance) |
Changes in gut microbiome affecting health and aging. | |||

|
Chronic inflammation (Inflammaging) |
Systemic inflammation contributing to aging and related diseases. | 2023[8] | ||

|
Altered intercellular communication | Changes in cellular communication leading to inflammation and tissue dysfunction. | 2013 | ||
Potential Hallmarks of Aging

|
Altered mechanical properties | Changes in cellular and extracellular structure affecting tissue function. | |

|
Splicing dysregulation
(Dysregulation in RNA splicing) |
Impaired RNA construction from DNA, leading to cellular dysfunction. | 2019[9] |
The Hallmarks in Detail
| Hallmark | Background | Manifests during normal aging | Experimentally increasing it accelerates aging | Experimentally amending it slows the normal aging process and increases healthy lifespan. | Associated human diseases |
|---|---|---|---|---|---|
Genomic instability
|
Damange in the DNA are formed mainly through oxidative stress and environmental factors.[10] A number of molecular processes work continuously to repair this damage.[11] | DNA damage accumulates over time[12] | Deficient DNA repair causes premature aging[13] | Increased DNA repair facilitates greater longevity[13] | |
Telomere attrition
|
Telomere attrition refers to the progressive shortening of telomeres, which are protective sequences at the ends of chromosomes. This occurs due to the inability of DNA polymerases to completely replicate the ends of linear DNA, and the absence of telomerase in most somatic cells. Shortened telomeres lead to cellular aging and reduced regenerative capacity, manifesting as replicative senescence or Hayflick limit[14]. Shelterins protect telomeres but may mask damage leading to persistent DNA damage and cellular stress[15]. Dysfunctions in telomere maintenance are linked to various age-related diseases[16]. | Telomere shortening is observed during normal aging in humans and mice[17]. | Excessive telomere attrition due to stress or genetic factors accelerates cellular aging and the onset of age-related pathologies[18]. | Maintaining telomere length through telomerase activation or shelterin integrity can delay aging and extend lifespan, as shown in mouse models and suggested by human epidemiological studies[19][20]. | Telomere shortening is associated with a variety of human diseases, including pulmonary fibrosis, dyskeratosis congenita, and aplastic anemia, often linked to deficiencies in telomerase or shelterin components[16]. |
Epigenetic alterations
|
Histone modifications are a type of epigenetic alteration that play a crucial role in regulating gene expression. Histones are proteins around which DNA is wrapped in eukaryotic cells, forming a structure known as a nucleosome. These modifications occur primarily at the tails of histone proteins and influence how tightly or loosely DNA is wound around the histones, affecting the accessibility of the DNA to various cellular machinery for processes like transcription, replication, and repair. | Chemical changes to histone proteins after they are formed can activate or silence gene expression and regulate the aging process.[21] | Sirtuins influence histone modifications:
|
Disorders in histone modification are linked with various aging-related conditions, implicating altered gene expression and protein function[26]. | |
| DNA methylation shift: DNA methylation is a biochemical process involving the addition of a methyl group to the DNA molecule, specifically to the cytosine or adenine DNA nucleotides. This process is a form of epigenetic modification, which means it can affect gene expression and function without changing the DNA sequence itself. | DNA methylation generally decreases with age in certain human and mouse tissues or cell cultures.[27][28][29][30] The loss of methylation in CD4+ T cells is proportional to age.[28] | No direct evidence yet. | No direct evidence yet. | Progeroid syndromes exhibit DNA methylation patterns similar to normal aging, suggesting a link with aging-related diseases[31][32]. | |
| Chromatin remodeling in the context of epigenetic alterations refers to the dynamic modification of the chromatin architecture to regulate access to genetic information in the DNA. Chromatin, which consists of DNA wrapped around histone proteins, can be altered or remodeled in various ways to either condense and silence gene regions or relax and activate them. This remodeling is a crucial mechanism for controlling gene expression, replication, repair, and other essential cellular processes. | Global canonical histone loss is regarded as a common feature of aging from yeast to humans.[33][34][35] | Flies with loss-of-function mutations in HP1α (a key chromosomal protein) have a shortened lifespan.[26] | Overexpression of HP1α extends longevity in flies and delays the muscular deterioration characteristic of old age.[26] | ||
| Transcriptional alterations refer to changes in the process by which genes are expressed into RNA, which can then be translated into proteins. These alterations can influence which genes are turned on or off, the timing of their activation, the amount of RNA produced, and the stability or processing of that RNA. | Aging leads to more transcriptional noise, meaning more random variations in gene activity[36], and also causes irregular production and processing of many mRNAs, the molecules that carry genetic information from DNA for protein creation[37][38]. Also miRNAs that is associated with the aging process is affected[39][40]. | Expression of several miRNAs increases longevity in Drosophila melanogaster and C. elegans[41][42][43]. | Loss of several miRNAs is associated with increase aging in Drosophila melanogaster and C. elegans[41][42][43]. | ||
| Loss of proteostasis | Loss of proteostasis refers to the disruption of the body's ability to regulate its proteins effectively, a process in which chaperones play a crucial role. This encompasses the processes of protein synthesis, folding, transport, and degradation. Chaperones, specialized proteins that assist in the proper folding and stabilization of other proteins, are essential in maintaining proteostasis. As we age, or in certain diseases, the balance of these processes can be disturbed, leading to the accumulation of misfolded or damaged proteins, and a decrease in the ability to produce and maintain healthy proteins. The decline in the efficiency or availability of chaperones contributes to this loss of proteostasis, exacerbating the accumulation of dysfunctional proteins and cellular stress. | Aging and some aging-related diseases are linked to impaired protein homeostasis or proteostasis.[44] | Mutant mice that that lack a certain helper chaperone from the heat-shock protein family show accelerated aging.[45] | Transgenic worms and flies overexpressing chaperones are long-lived[46][47] | Chronic expression of unfolded, misfolded or aggregated proteins contributes to the development of some age-related pathologies, such as Alzheimer’s disease, Parkinson’s disease and cataracts[44]. |
| Disabled macroautophagy | Disabled macroautophagy, often referred as impaired or dysfunctional autophagy, is a condition where the cellular process of autophagy—specifically the macroautophagy pathway—is disrupted or less effective. Autophagy is a critical cellular process for degrading and recycling damaged organelles, misfolded proteins, and other cellular debris. Macroautophagy involves the engulfment of these unwanted materials into vesicles called autophagosomes, which then fuse with lysosomes where the contents are degraded and recycled. When macroautophagy is disabled or impaired, cells accumulate damaged proteins and organelles, leading to cellular dysfunction and contributing to various diseases, particularly those related to aging and neurodegeneration. This loss of a crucial cellular "cleanup" mechanism can result in increased oxidative stress, disrupted cellular homeostasis, and an acceleration of the aging process.
While originally considered under hallmark altered proteostasis, autophagy regulates a number of other hallmarks of ageing such as DNA repair and nutrient sensing/metabolism[48], and hence it was proposed to be categorized as an integrative hallmark. |
Compromised autophagy is observed in numerous ageing conditions including neurodegeneration and immunosenescence[49][50]. | Activation of autophagy can increase mouse lifespan[51], and even improve immune response to vaccination in older humans by overcoming immunosenescence[52]. | ||
| Deregulated nutrient sensing | Deregulated nutrient sensing refers to the body's declining ability to properly manage and respond to nutrients, such as fats, sugars, and proteins, as we get older. Normally, our bodies have finely tuned systems that detect when we eat and use these nutrients efficiently for energy and repair. However, with age, these systems start to malfunction. This means our body might not handle sugars well, leading to conditions like diabetes, or it might struggle with managing fats, leading to issues like high cholesterol. Essentially, Deregulated Nutrient Sensing is when our body's 'nutrient management system' becomes less efficient and accurate with age, leading to various metabolic and health problems. | Deregulated nutrient sensing ability takes place upon aging.[53] | The significance of nutrient sensing throughout the aging process has been first established in the prominent observation that decreased food intake in rats prolongs lifespan relative to ad libitum fed controls.[54] | ||
Mitochondrial dysfunction
|
|||||
| Cellular senescence | |||||
Stem cell exhaustion
|
|||||
| Dysbiosis (Microbiome disturbance) |
|||||
| Chronic inflammation (Inflammaging) |
|||||
Altered intercellular communication
|
Correlation to Age-Related Diseases
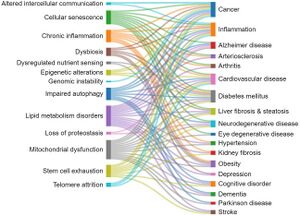
This section explores the correlations between the aging hallmarks and the age-related diseases, as reflected in the number of documents in the CAS Content Collection. Generally, cellular senescence, mitochondrial dysfunction, lipid metabolism disorders, and inflammaging appear as related to multiple pathologies.[2]
Some particular correlations are noteworthy:
- There is a strong correlation between documents related to cellular senescence and cancer, according to the CAS Content Collection. Cellular senescence is a state of a cell cycle arrest, so the entry of cells into senescence can act as a barrier to tumorigenesis thus being of special interest for anticancer therapies. It has been demonstrated however that, in certain conditions, malignant and nonmalignant senescent cells can develop protumorigenic properties and eventually trigger tumor relapse, evidencing contrasting roles of senescent cells in cancer still remaining to be explored.[55][56][57]
- The strongest correlation between diabetes mellitus and aging hallmarks is with the lipid metabolism disorders, according to the CAS Content Collection documents number. Glucose and lipid metabolism are correlated in multiple ways.[58] One of the notable manifestations of this correlation is diabetic dyslipidemia, with both being well established cardiovascular risk factors. The link between glucose and lipid metabolism is in fact rather complex with both lipids and glucose playing important roles in energy metabolism.[58][59][60]
- Hypertension–lipid metabolism disorders correlation: It has been reported that both hypertension and aging are associated with higher lipid peroxidation.[61] Aging is additionally associated with an increase in lipid peroxidation in cardiac muscle.[62]
- Inflammation–cellular senescence correlation: Aging is characterized by systemic chronic inflammation, linked to cellular senescence, immunosenescence, and age-related organ dysfunction. Senescence-associated secretory phenotype (SASP) factors secreted by senescent cells promote chronic inflammation. Meanwhile, chronic inflammation accelerates the senescence of immune cells, resulting in an inability to clear inflammatory factors, which creates a malicious cycle of inflammation and senescence.
- Altogether, there is significant correlation between cellular senescence and the majority of age-related diseases.[63] The disadvantages of senescence seem to be in, first, causing a loss of tissue-repair capacity because of cell cycle arrest in progenitor cells and, second, in producing proinflammatory molecules in the senescence-associated secretory phenotype (SASP). Substantial pool of information about senescence in cells has been acquired recently; however, it is still poorly understood.
- Cognitive impairment–mitochondrial dysfunction correlation: The brain profoundly depends on mitochondria to produce energy, in order to maintain essential bodily functions. Upon aging, damaged mitochondria accumulate. They produce insufficient ATP and excessive ROS. It has been recently reported that mitochondria at dysfunctional synapses do not meet the energetic need and potentially trigger age-related cognitive impairment.[64][65]
- Alzheimer disease–mitochondrial dysfunction correlation: Alzheimer’s disease is the most frequent source of age-related neurodegeneration and cognitive impairment. A growing body of evidence implicates mitochondrial dysfunction as a common pathogenic mechanism involved in many of the features of the Alzheimer’s patients brain, such as formation of amyloid plaques and neurofibrillary tangles.[66]
- Altogether, there is significant correlation between mitochondrial dysfunction and the majority of age-related diseases including diabetes, inflammation, obesity, neurodegenerative disorders, cardiovascular diseases, and cancer.[67] Mitochondria are vital in regulation of energy and metabolic homeostasis. Proper mitochondrial functions, including cellular energy production and control of oxidative stress, are in strong relation with the accurate performance of brain, cognition, and the overall health.[68]
- Liver fibrosis–lipid metabolic disorders correlation: Liver plays a key role in lipid metabolism; therefore alterations in hepatic lipid metabolism can be a factor in development of chronic liver disease. Furthermore, chronic liver disease can impact hepatic lipid metabolism causing alterations in circulating lipid levels contributing to dyslipidemia.[69] Likewise, the liver plays an essential role in lipid metabolism, certain steps of lipid synthesis, and transport. Therefore, abnormal lipid profiles and liver dysfunctions are expectedly closely correlated.[70]
- Altogether, there is significant correlation between lipid metabolic disorders and the majority of age-related diseases.[71] Upon aging, body fat builds up with changes in the lipid metabolism. Considering lipid metabolism, excess body fat with enhanced lipotoxicity triggers various age-related diseases, including cardiovascular disease, cancer, arthritis, diabetes, and Alzheimer’s disease. Progress in lipidomic techniques has identified alterations in lipid profiles associated with aging. Lipid accumulation and impaired fatty acid processing are associated with pathophysiological aging phenotypes. Although it is still not well-known how lipid metabolism is regulated upon aging, data suggest a dynamic role for lipid metabolism in signaling and gene expression regulation.[71][72]
History
- 2013 The scientific journal Cell published the article "The Hallmarks of Aging", that was translated to several languages and determined the directions of many studies.[1]
- 2022 It was proposed to expand the list of the nine hallmarks of aging with five more.[5][73]
- 2023 In a paywalled review, the authors of a heavily cited paper on the hallmarks of aging update the set of proposed hallmarks after a decade.[6] A review with overlapping authors merge or link various hallmarks of cancer with those of aging.[74]
Further Reading
- 2013, The hallmarks of aging [1]
- 2023, Aging Hallmarks and Progression and Age-Related Diseases: A Landscape View of Research Advancement [2]
- 2023, Hallmarks of aging: An expanding universe [6]
See Also
- Age-Related Diseases
- Wikipedia - Hallmarks of aging
Todo
- 2023, Chronic inflammation and the hallmarks of aging [8]
- 2022, Biological mechanisms of aging predict age-related disease co-occurrence in patients [3]
- https://doi.org/10.1038/s41587-023-02024-y
- 2023, Meta-hallmarks of aging and cancer [74]
References
- ↑ Jump up to: 1.0 1.1 1.2 1.3 1.4 1.5 López-Otín C et al.: The hallmarks of aging. Cell 2013. (PMID 23746838) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 2.0 2.1 2.2 2.3 Tenchov R et al.: Aging Hallmarks and Progression and Age-Related Diseases: A Landscape View of Research Advancement. ACS Chem Neurosci 2023. (PMID 38095562) [PubMed] [DOI] Abstract
- ↑ Jump up to: 3.0 3.1 Fraser HC et al.: Biological mechanisms of aging predict age-related disease co-occurrence in patients. Aging Cell 2022. (PMID 35259281) [PubMed] [DOI] [Full text] Abstract
- ↑ Gems D & de Magalhães JP: The hoverfly and the wasp: A critique of the hallmarks of aging as a paradigm. Ageing Res Rev 2021. (PMID 34271186) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 5.0 5.1 Schmauck-Medina T et al.: New hallmarks of ageing: a 2022 Copenhagen ageing meeting summary. Aging (Albany NY) 2022. (PMID 36040386) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 6.0 6.1 6.2 López-Otín C et al.: Hallmarks of aging: An expanding universe. Cell 2023. (PMID 36599349) [PubMed] [DOI] Abstract
- ↑ Kaushik S et al.: Autophagy and the hallmarks of aging. Ageing Res Rev 2021. (PMID 34563704) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 8.0 8.1 Baechle JJ et al.: Chronic inflammation and the hallmarks of aging. Mol Metab 2023. (PMID 37329949) [PubMed] [DOI] [Full text] Abstract
- ↑ Bhadra M et al.: Alternative splicing in aging and longevity. Hum Genet 2020. (PMID 31834493) [PubMed] [DOI] [Full text] Abstract
- ↑ De Bont R & van Larebeke N: Endogenous DNA damage in humans: a review of quantitative data. Mutagenesis 2004. (PMID 15123782) [PubMed] [DOI] Abstract
- ↑ de Duve C: The onset of selection. Nature 2005. (PMID 15703726) [PubMed] [DOI]
- ↑ Vijg J & Suh Y: Genome instability and aging. Annu Rev Physiol 2013. (PMID 23398157) [PubMed] [DOI] Abstract
- ↑ Jump up to: 13.0 13.1 Hoeijmakers JH: DNA damage, aging, and cancer. N Engl J Med 2009. (PMID 19812404) [PubMed] [DOI]
- ↑ Blackburn EH et al.: Telomeres and telomerase: the path from maize, Tetrahymena and yeast to human cancer and aging. Nat Med 2006. (PMID 17024208) [PubMed] [DOI]
- ↑ Palm W & de Lange T: How shelterin protects mammalian telomeres. Annu Rev Genet 2008. (PMID 18680434) [PubMed] [DOI] Abstract
- ↑ Jump up to: 16.0 16.1 Armanios M & Blackburn EH: The telomere syndromes. Nat Rev Genet 2012. (PMID 22965356) [PubMed] [DOI] [Full text] Abstract
- ↑ Blasco MA: Telomere length, stem cells and aging. Nat Chem Biol 2007. (PMID 17876321) [PubMed] [DOI] Abstract
- ↑ Fumagalli M et al.: Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nat Cell Biol 2012. (PMID 22426077) [PubMed] [DOI] [Full text] Abstract
- ↑ Jaskelioff M et al.: Telomerase reactivation reverses tissue degeneration in aged telomerase-deficient mice. Nature 2011. (PMID 21113150) [PubMed] [DOI] [Full text] Abstract
- ↑ Bernardes de Jesus B et al.: Telomerase gene therapy in adult and old mice delays aging and increases longevity without increasing cancer. EMBO Mol Med 2012. (PMID 22585399) [PubMed] [DOI] [Full text] Abstract
- ↑ Kouzarides T: Chromatin modifications and their function. Cell 2007. (PMID 17320507) [PubMed] [DOI] Abstract
- ↑ Mostoslavsky R et al.: Genomic instability and aging-like phenotype in the absence of mammalian SIRT6. Cell 2006. (PMID 16439206) [PubMed] [DOI] Abstract
- ↑ Herranz D et al.: Sirt1 improves healthy ageing and protects from metabolic syndrome-associated cancer. Nat Commun 2010. (PMID 20975665) [PubMed] [DOI] [Full text] Abstract
- ↑ Brown K et al.: SIRT3 reverses aging-associated degeneration. Cell Rep 2013. (PMID 23375372) [PubMed] [DOI] [Full text] Abstract
- ↑ Kanfi Y et al.: The sirtuin SIRT6 regulates lifespan in male mice. Nature 2012. (PMID 22367546) [PubMed] [DOI] Abstract
- ↑ Jump up to: 26.0 26.1 26.2 Larson K et al.: Heterochromatin formation promotes longevity and represses ribosomal RNA synthesis. PLoS Genet 2012. (PMID 22291607) [PubMed] [DOI] [Full text] Abstract
- ↑ Wilson VL et al.: Genomic 5-methyldeoxycytidine decreases with age. J Biol Chem 1987. (PMID 3611071) [PubMed] Abstract
- ↑ Jump up to: 28.0 28.1 Heyn H et al.: Distinct DNA methylomes of newborns and centenarians. Proc Natl Acad Sci U S A 2012. (PMID 22689993) [PubMed] [DOI] [Full text] Abstract
- ↑ Zhang J et al.: Highly enriched BEND3 prevents the premature activation of bivalent genes during differentiation. Science 2022. (PMID 35143257) [PubMed] [DOI] Abstract
- ↑ Seale K et al.: Making sense of the ageing methylome. Nat Rev Genet 2022. (PMID 35501397) [PubMed] [DOI] [Full text] Abstract
- ↑ Osorio FG et al.: Nuclear envelope alterations generate an aging-like epigenetic pattern in mice deficient in Zmpste24 metalloprotease. Aging Cell 2010. (PMID 20961378) [PubMed] [DOI] Abstract
- ↑ Shumaker DK et al.: Mutant nuclear lamin A leads to progressive alterations of epigenetic control in premature aging. Proc Natl Acad Sci U S A 2006. (PMID 16738054) [PubMed] [DOI] [Full text] Abstract
- ↑ Dang W et al.: Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 2009. (PMID 19516333) [PubMed] [DOI] [Full text] Abstract
- ↑ Hu Z et al.: Nucleosome loss leads to global transcriptional up-regulation and genomic instability during yeast aging. Genes Dev 2014. (PMID 24532716) [PubMed] [DOI] [Full text] Abstract
- ↑ O'Sullivan RJ et al.: Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat Struct Mol Biol 2010. (PMID 20890289) [PubMed] [DOI] [Full text] Abstract
- ↑ Bahar R et al.: Increased cell-to-cell variation in gene expression in ageing mouse heart. Nature 2006. (PMID 16791200) [PubMed] [DOI] Abstract
- ↑ Harries LW et al.: Human aging is characterized by focused changes in gene expression and deregulation of alternative splicing. Aging Cell 2011. (PMID 21668623) [PubMed] [DOI] [Full text] Abstract
- ↑ Nicholas A et al.: Age-related gene-specific changes of A-to-I mRNA editing in the human brain. Mech Ageing Dev 2010. (PMID 20538013) [PubMed] [DOI] [Full text] Abstract
- ↑ Toledano H et al.: The let-7-Imp axis regulates ageing of the Drosophila testis stem-cell niche. Nature 2012. (PMID 22660319) [PubMed] [DOI] [Full text] Abstract
- ↑ Ugalde AP et al.: Micromanaging aging with miRNAs: new messages from the nuclear envelope. Nucleus 2011. (PMID 22064465) [PubMed] [DOI] Abstract
- ↑ Jump up to: 41.0 41.1 Liu N et al.: The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature 2012. (PMID 22343898) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 42.0 42.1 Shen Y et al.: A steroid receptor-microRNA switch regulates life span in response to signals from the gonad. Science 2012. (PMID 23239738) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 43.0 43.1 Smith-Vikos T & Slack FJ: MicroRNAs and their roles in aging. J Cell Sci 2012. (PMID 22294612) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 44.0 44.1 Powers ET et al.: Biological and chemical approaches to diseases of proteostasis deficiency. Annu Rev Biochem 2009. (PMID 19298183) [PubMed] [DOI] Abstract
- ↑ Min JN et al.: CHIP deficiency decreases longevity, with accelerated aging phenotypes accompanied by altered protein quality control. Mol Cell Biol 2008. (PMID 18411298) [PubMed] [DOI] [Full text] Abstract
- ↑ Morrow G et al.: Overexpression of the small mitochondrial Hsp22 extends Drosophila life span and increases resistance to oxidative stress. FASEB J 2004. (PMID 14734639) [PubMed] [DOI] Abstract
- ↑ Walker GA & Lithgow GJ: Lifespan extension in C. elegans by a molecular chaperone dependent upon insulin-like signals. Aging Cell 2003. (PMID 12882326) [PubMed] [DOI] Abstract
- ↑ Kaushik S & Cuervo AM: The coming of age of chaperone-mediated autophagy. Nat Rev Mol Cell Biol 2018. (PMID 29626215) [PubMed] [DOI] [Full text] Abstract
- ↑ Wong SQ et al.: Autophagy in aging and longevity. Hum Genet 2020. (PMID 31144030) [PubMed] [DOI] [Full text] Abstract
- ↑ Aman Y et al.: Autophagy in healthy aging and disease. Nat Aging 2021. (PMID 34901876) [PubMed] [DOI] [Full text] Abstract
- ↑ Fernández ÁF et al.: Disruption of the beclin 1-BCL2 autophagy regulatory complex promotes longevity in mice. Nature 2018. (PMID 29849149) [PubMed] [DOI] [Full text] Abstract
- ↑ Alsaleh G et al.: Autophagy in T cells from aged donors is maintained by spermidine and correlates with function and vaccine responses. Elife 2020. (PMID 33317695) [PubMed] [DOI] [Full text] Abstract
- ↑ Falcón P et al.: Nutrient Sensing and Redox Balance: GCN2 as a New Integrator in Aging. Oxid Med Cell Longev 2019. (PMID 31249645) [PubMed] [DOI] [Full text] Abstract
- ↑ McCay CM et al.: The effect of retarded growth upon the length of life span and upon the ultimate body size. 1935. Nutrition 1989. (PMID 2520283) [PubMed]
- ↑ Schmitt CA et al.: Senescence and cancer - role and therapeutic opportunities. Nat Rev Clin Oncol 2022. (PMID 36045302) [PubMed] [DOI] [Full text] Abstract
- ↑ Santos-de-Frutos K & Djouder N: When dormancy fuels tumour relapse. Commun Biol 2021. (PMID 34135460) [PubMed] [DOI] [Full text] Abstract
- ↑ Yang J et al.: The Paradoxical Role of Cellular Senescence in Cancer. Front Cell Dev Biol 2021. (PMID 34458273) [PubMed] [DOI] [Full text] Abstract
- ↑ Jump up to: 58.0 58.1 Parhofer KG: Interaction between Glucose and Lipid Metabolism: More than Diabetic Dyslipidemia. Diabetes Metab J 2015. (PMID 26566492) [PubMed] [DOI] [Full text] Abstract
- ↑ Athyros VG et al.: Diabetes and lipid metabolism. Hormones (Athens) 2018. (PMID 29858856) [PubMed] [DOI] Abstract
- ↑ Savage DB et al.: Disordered lipid metabolism and the pathogenesis of insulin resistance. Physiol Rev 2007. (PMID 17429039) [PubMed] [DOI] [Full text] Abstract
- ↑ Yavuzer H et al.: Biomarkers of lipid peroxidation related to hypertension in aging. Hypertens Res 2016. (PMID 26763852) [PubMed] [DOI] Abstract
- ↑ Miró O et al.: Aging is associated with increased lipid peroxidation in human hearts, but not with mitochondrial respiratory chain enzyme defects. Cardiovasc Res 2000. (PMID 10963736) [PubMed] [DOI] Abstract
- ↑ Childs BG et al.: Cellular senescence in aging and age-related disease: from mechanisms to therapy. Nat Med 2015. (PMID 26646499) [PubMed] [DOI] [Full text] Abstract
- ↑ Mitochondrial Dysfunction May Be a Cause of Age-Related Cognitive Impairment. https://www.genengnews.com/news/mitochondrial-dysfunction-may-be-a-cause-of-age-related-cognitive-impairment/#:~:text=During%20aging%2C%20damaged%20mitochondria%20that,cause%20age%2Drelated%20cognitive%20impairment. (accessed Jul 21, 2023).
- ↑ Glavis-Bloom C et al.: Violation of the ultrastructural size principle in the dorsolateral prefrontal cortex underlies working memory impairment in the aged common marmoset (Callithrix jacchus). Front Aging Neurosci 2023. (PMID 37122384) [PubMed] [DOI] [Full text] Abstract
- ↑ Sharma C et al.: Mitochondrial Dysfunction as a Driver of Cognitive Impairment in Alzheimer's Disease. Int J Mol Sci 2021. (PMID 34063708) [PubMed] [DOI] [Full text] Abstract
- ↑ Srivastava S: The Mitochondrial Basis of Aging and Age-Related Disorders. Genes (Basel) 2017. (PMID 29257072) [PubMed] [DOI] [Full text] Abstract
- ↑ Geary DC: Mitochondrial Functioning and the Relations among Health, Cognition, and Aging: Where Cell Biology Meets Cognitive Science. Int J Mol Sci 2021. (PMID 33808109) [PubMed] [DOI] [Full text] Abstract
- ↑ Arvind A.; Osganian S. A.; Cohen D. E.E.; C K.. Lipid and Lipoprotein Metabolism in Liver Disease. In Endotext [Internet]; Feingold K. R., Anawalt B., Blackman M. R., Eds.; MDText.com, Inc.: South Dartmouth, 2019. [Google Scholar]
- ↑ Ghadir MR et al.: The relationship between lipid profile and severity of liver damage in cirrhotic patients. Hepat Mon 2010. (PMID 22312394) [PubMed] [Full text] Abstract
- ↑ Jump up to: 71.0 71.1 Chung KW: Advances in Understanding of the Role of Lipid Metabolism in Aging. Cells 2021. (PMID 33924316) [PubMed] [DOI] [Full text] Abstract
- ↑ Johnson AA & Stolzing A: The role of lipid metabolism in aging, lifespan regulation, and age-related disease. Aging Cell 2019. (PMID 31560163) [PubMed] [DOI] [Full text] Abstract
- ↑ Researchers Propose Five New Hallmarks of Aging, https://www.lifespan.io/news/researchers-propose-five-new-hallmarks-of-aging/
- ↑ Jump up to: 74.0 74.1 López-Otín C et al.: Meta-hallmarks of aging and cancer. Cell Metab 2023. (PMID 36599298) [PubMed] [DOI] Abstract
